[ad_1]
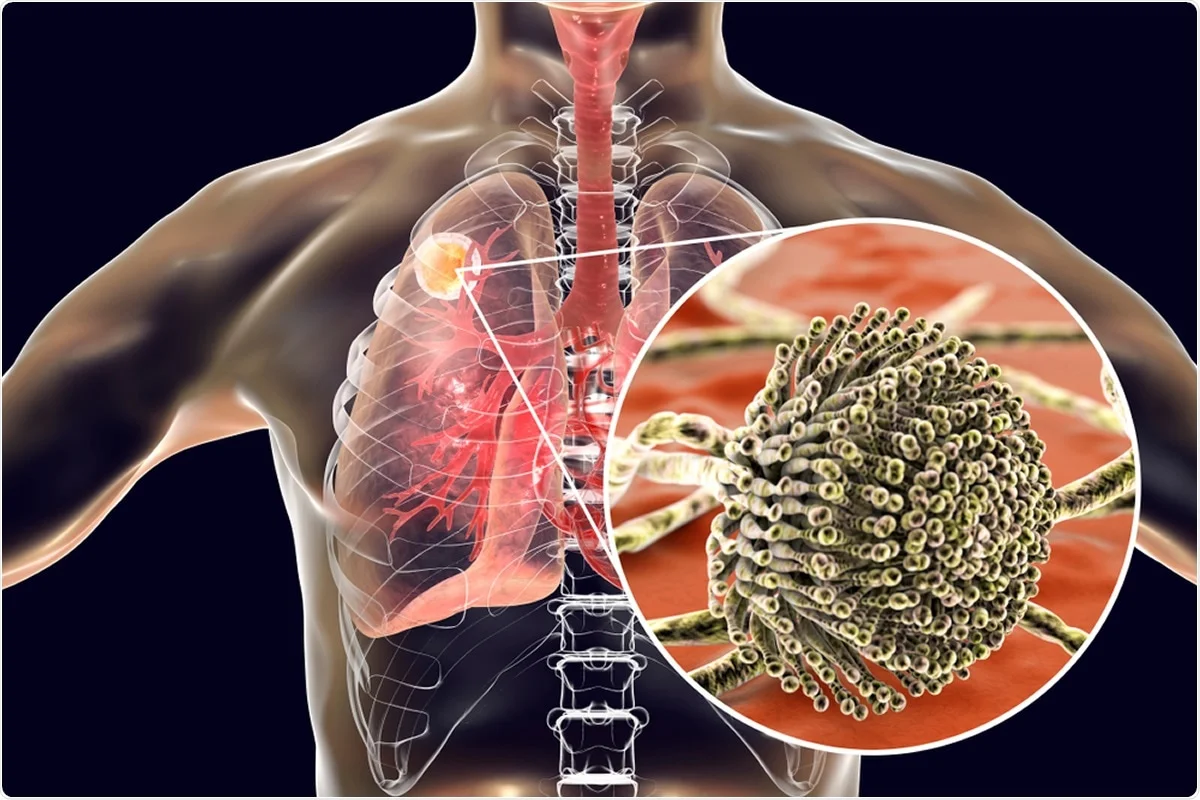
Caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), coronavirus disease 2019 (COVID-19) is primarily a lung disease, and a large percentage of patients with severe cases also have a bacterial infection or secondary fungal. In fact, about 25% of patients with severe acute respiratory distress syndrome (ARDS), and about 40% of patients with severe COVID-19 pneumonia, also show signs of fungal infection. Aspergillus fumigatus in the lungs.
A new study published on the prepress server bioRxiv* in November 2020 discusses the characteristics of COVID-19 associated pulmonary aspergillosis (CAPA).

This fungus causes invasive lung infection, following inhalation of asexual spores. There are over 250,000 cases of aspergillosis every year and these are associated with many deaths. This fungus is able to grow at the temperature of the human body, is resistant to oxidation and rapidly develops resistance to antifungals.
The current study aims to understand whether CAPA isolates are different from others A. fumigatus strains with regard to their genetic composition or phenotypic expression. The researchers looked at four of these isolates from four patients admitted to an intensive care unit (ICU) with ARDS due to critical COVID-19.

Inhalation of Aspergillus spores can cause fungal infections. Inhaling Aspergillus spores from the environment can reach the lung and then grow vegetatively and spread to other parts of the body.
CAPA isolates belong to A. fumigatus
The study found that all patients had a A. fumigatus tensions. The CAPA isolates were also closely related and appear to share a common ancestor. CAPA isolates are also similar to the reference strains Af293 and CEA17.
All 206 genes affected by virulence in A. fumigatus were present in CAPA isolates with no change in copy number, but exhibited single nucleotide polymorphisms (SNPs) and indel mutations, which correlated with the change in expected effect.
LOF mutations in virulence genes
The numerous polymorphisms observed in CAPA isolates resulted in early termination or frameshift mutations, both of which may indicate loss of function (LOF) in NRPS8, which is a peptide synthase gene responsible for the production of an unknown secondary metabolite. Such mutations in these genes were found to increase virulence in a mouse experiment.
Many of the (possible) LOF mutations also occurred between genes that had null mutants linked to decreased virulence, such as premature termination of the PPTA gene. Deletion mutants of the latter were found to cause less virulence in mice. The researchers pointed out that the monophyletic relationship between the four isolates means that the shared SNPs observed in PPTA across all CAPA isolates can be traced to a mutational event in the genome of the last common ancestral strain.
Specific polymorphisms of the isolate
However, several isolates also showed specific polymorphisms that affect virulence. One of these mutations affects CYP5081A1, a cytochrome enzyme in CAPA isolates B and C, and one in the FLEA gene in isolate D. The FLEA null mutant is associated with more severe pneumonia and with a higher likelihood of invasive aspergillosis in mouse studies . This is because this gene binds to macrophages and its presence is necessary for the recognition of the host by the virus, as well as for the killing of macrophages.
Biosynthetic gene clusters
Again, all CAPA isolates had clusters of biosynthetic (BGC) genes that synthesize toxic or modulatory metabolites, such as gliotoxin, which was produced by all isolates, or fumagillin, which is known to inhibit neutrophil function. . All CAPA isolates showed BGCs of equivalent class and similar number.
The largest number of LOF mutations among genes associated with higher virulence were found in the CAPA isolate D. This conclusion is supported by the pairwise comparison of pathogenicity between isolates, which showed this to be the most virulent, with isolate C also showing markedly higher virulence than the reference strains.
The pathogenicity profile of the CAPA isolates closely resembled that of the reference strains. They were more sensitive to the chemical white calcofluor than the reference strains, but did not show significant differences in susceptibility to antifungals.
Conclusions
The researchers concluded, therefore, that the CAPA isolates are closely related and contain genes for virulence with almost no attenuation. Furthermore, these genes are expressed as one would expect from this fungus. However, there were genetic and phenotypic differences between the virulence determinants of the different isolates.
Isolate D appears to be more virulent than other strains (both CAPA isolates and reference strains) as assessed in an invertebrate infection model.
“We hypothesize that this difference is due to the fact that isolate D contains more putative LOF mutations in the genetic virulence determinants whose null mutants are known to increase virulence compared to other strains,“The researchers said.
*Important Notice
bioRxiv publishes preliminary scientific reports that are not peer-reviewed and therefore should not be considered conclusive, guide clinical practice / health-related behavior, or treated as consolidated information.
.
[ad_2]
Source link