[ad_1]
In a recent medRxiv * preprint paper, the researchers of the Health Center of the University of Connecticut in the United States reveal their method CRISPR (WS-CRISPR) with digital hot start for rigorous quantitative and sensitive detection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV -2) in clinical samples from patients with coronavirus disease (COVID-19).
Since an effective antiviral treatment and COVID-19 vaccines fully validated are not available so far in our battle against COVID-19, the precise and accurate quantitation of SARS-CoV-2 virus plays a key role in the assessment of infectivity parameters and control of pandemic in progress.
In order to quantify SARS-CoV-2, probe-based reverse transcription polymerase chain reaction (RT-PCR) is currently used as a gold standard technique due to its relatively high sensitivity and specificity; however, it depends on expensive instruments and well-designed probes not suitable for small clinics or community health settings.
Likewise, isothermal amplification methods are effectively intended for qualitative detection purposes or are frequently exposed to unwanted non-specific amplification or false positive signals.
This is where CRISPR-Cas based nucleic acid detection comes into play. However, the question is whether it can be modified to allow for accurate quantification without unwanted premature amplification of the target and subsequent overestimation if digital sensing is used.
A US research team, led by Ding dott.Xiong of the Biomedical Engineering Department at the University of Connecticut Health Center in Farmington, described a WS-digital CRISPR essay that promises a valid and reliable quantification of nucleic SARS- CoV-2 acids in clinical specimens.

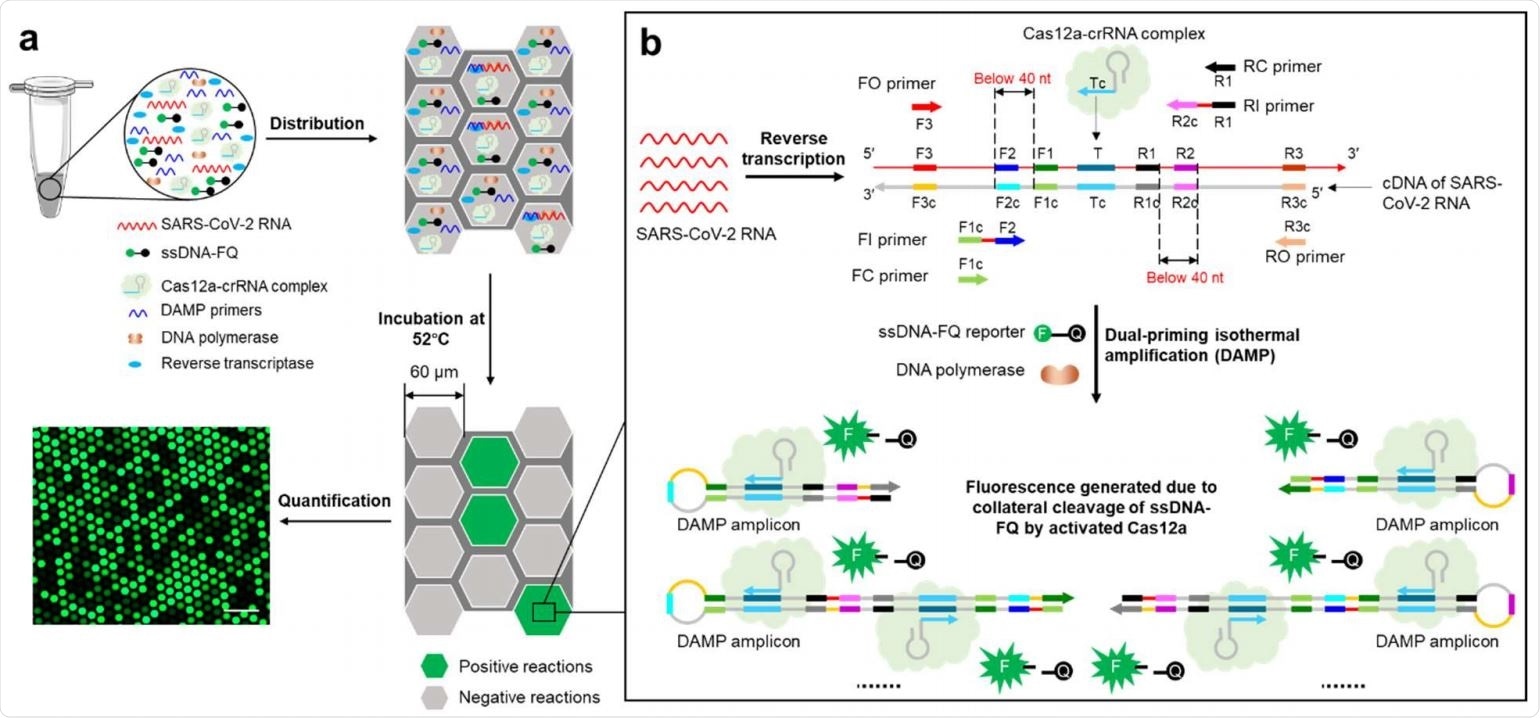
Digital WS-CRISPR Assay Overview. a, The WS-CRISPR reaction mix to a jar is first prepared in a tube. After deployment in the QuantStudio 3D digital chip, over ten thousand sub-nanoliter (~ 0.7 nL) micro-reactions are isolated in microwells. When incubated at 52 ° C, each micro-reaction with the SARS-CoV-2 RNA target undertakes the WS-CRISPR reaction and generates a strong green fluorescence (positive spots), while not in those without targets (negative spots). The scale bar is 300 μm. Through the detection and counting of positive microreactions (or spots), SARS-CoV-2 RNA can be quantified based on the proportion of positive spots. b, Principle of operation of the one-cup WS-CRISPR test for SARS-CoV-2 detection. The WSCRISPR reaction mix contains the Cas12a-crRNA complex, six DAMP primers (two external primers of FO and RO, two internal primers of FI and RI and two competition primers of FC and RC), ssDNA-FQ reporter, SuperScript reverse transcriptase IV, Bst DNA polymerase and SARS-CoV-2 RNA in a single vessel format.
Constitutive elements of the new method
In this exciting scientific endeavor, researchers took advantage of a hot start CRISPR reaction to an inaugural pot, which combines low-temperature double reverse transcription-mediated isothermal amplification (RT-DAMP) and CRISPR-Cas12a-based detection. .
More specifically, to couple these two different reaction systems in a vessel, pyrophosphatase was added to maintain a constant concentration of magnesium ions by degrading the magnesium pyrophosphate byproduct.
Furthermore, phosphorothioated internal primers were used in this study to allow for effective isothermal amplification of reverse transcription at rather low temperatures (such as 52 ° C).
Finally, by dividing the reaction mixture in one-pot microreactions sub-nanoliter using digital 3D QuantStudio chip, this research group has developed a digital CRISPR test that allows the sensitive and reliable quantification of SARS-CoV-2.
Moving the CRISPR diagnostic milestone
Compared to the CRISPR-based nucleic acid techniques reported earlier, this digital WS-CRISPR assay offers several outstanding advantages. First of all, this is an inaugural demonstration of a one-pot CRISPR assay combining Bacillus stearothermophilus (Bst) Isothermal amplification of reverse transcription based on DNA polymerase with CRISPR-Cas12a detection, without the need for a higher reaction temperature and longer nucleic acids.
In addition, the method is typically initiated at an elevated temperature (above 50 ° C), addressing the problem unwanted premature amplification of the target in the digital detection, in addition to displaying a sensitivity 10 times higher and high specificity of detection compared to bulk based on assay format tubes.
By targeting the SARS-CoV-2 nucleoprotein gene, it is possible to quantify up to 5 copies of SARS-CoV-2 RNA per microliter in the chip with the use of the digital WS-CRISPR assay. Additionally, quantification can take place in clinical samples, allowing for assessment of COVID-19 infectivity and the effectiveness of antiviral drugs.
Finally, the proposed WS-CRISPR digital assay shows a high level of tolerance to inhibitors and can be used to directly detect the virus in raw saliva samples, circumventing the need for RNA extraction. This, in turn, will facilitate COVID-19 diagnosis and reduce the risk of infection in healthcare professionals.
A look to the future
“As the first clinically validated digital CRISPR test, our digital WS-CRISPR test provides reliable, sensitive and direct quantitative SARS-CoV-2 detection,” the study researchers point out. medRxiv prepress paper. “It opens up a new exploration for quantitative detection of CRISPR-based nucleic acid,” they add.

Direct detection of SARS-CoV-2 in raw saliva samples by digital WS-CRISPR assay. a, Workflow for direct SARS-CoV-2 testing in spiked saliva samples using WS-CRISPR digital assay. b, Chip endpoint fluorescence micrographs for direct detection of SARS-CoV-2 virus spiked in saliva samples. Saliva samples 1-5, samples with 10%, 5%, 2.5%, 1% and 0% heat inactivated SARS-CoV-2 virus. Each micrograph is a representative of six distinct regions taken to cover approximately 2809 microreactions. PC, SARS-CoV-2 positive control sample. NC, negative control sample for SARS-CoV-2. NTC, model-free control. The scale bars are 300 μm.
Despite the above benefits, a new digital chip needs to be further explored and validated for the WS-CRISPR digital assay in the future, requiring further clinical validation steps.
Furthermore, although this digital WS-CRISPR assay uses relatively expensive fluorescence microscopy for imaging purposes, smartphone-based handheld fluorescence microscopy could become the alternative option towards quantitative in situ detection in the near future.
*Important Notice
medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be considered conclusive, guide clinical practice / health-related behavior, or treated as consolidated information.
.
[ad_2]
Source link